Simple Electron Microscopy Primer

In order to reconstruct a 3D molecule from it's 2D projections, it is often necessary to use several thousand experimental projections (particles). For small sample sizes (e.g. less than two thousand), it is straightforward to select (pick) the particles from several micrographs. Using a computer program such as SPIDER/WEB, the untilted and tilted images (for random conical tilt) of the same micrograph are displayed side by side. Several particle pairs are then picked by hand. From their geometrical relationships, this allows the program to determine the angular relationships between the pairs (the exact tilt between micrographs and any rotation of the micrographs perpendicular to the tilt).
After the angles have been determined, usually only the untilted particle need to be selected by the user, as the location of the tilted particle is calculated automatically using the previously calculated angles. Of course, it is wise from time to time to recalculate the angles, however, as drift can occur from one area to another of the micrograph. This procedure is repeated for as many micrograph pairs as there are. In reconstructions that do not need tilt pairs (as when a reference is used, or when using the common lines method), picking particles is much easier, since any suitable particle can be used (there is no need for a matching tilted pair).

This procedure, of course, can become tedious after several dozen micrographs have been scanned. There is no real easy way to automatically pick all particles in tilt pairs, but when only single particles are needed, efficient methods do exist. Automatic particle picking procedures can use peak finding algorithms (of bright particles on a dark background) to select particles and avoid noise. A rough model of a particle is generated, and then that model is 'dragged' across the micrograph. When a good correlation between model and micrograph is made, this indicates that a possible particle has been found and the area is selected by the program and windowed out into its own file.
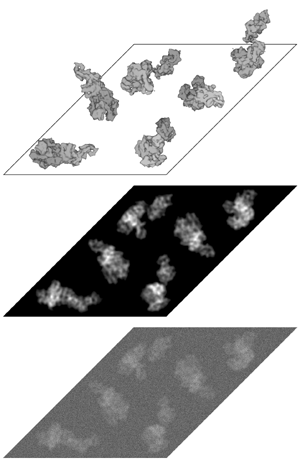
Another common method of automatic picking uses two steps: training and selection. In the training step, a small amount of particles are picked by the computer, and then the operator manually separates them into particles, junk, and background noise. The computer uses this information to design a discriminant function and then proceeds to pick the particles. With enough training, good programs should be able to differentiate between the various possibilities. Sometimes, too, the procedures can be calibrated to pick particles that are more than a set distance apart in order to avoid overlaps and picking the same particle twice. Unfortunately, these procedures generally only work on sample preparations that are homogenous. Large particle aggregates, as well as other artefacts in the scanned image, can confuse the picking. For these reasons, there is often a quality control step where a person manually scans the picked particles to ensure a low level of false positives. Nevertheless, once all the particle coordinates have been determined for the micrographs, the particles are 'windowed' out into their own individual files, where they can be manipulated in a more efficient manner. Some programs simplify the task of keeping track of so many particles by 'stacking' them into one file. In a stack, all the images are concatenated into a continuous data stream, where it is easier (and quicker) for the program to access.